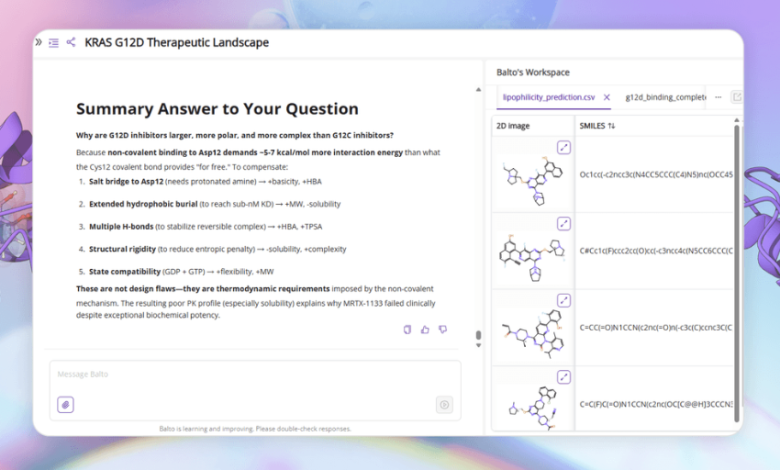
Scroll any life sciences feed and you’ll see the same headline wearing a different hat: “AI will revolutionize drug discovery.”
Sometimes it will. But most teams don’t lose years because they lack a moonshot. They lose years because brilliant scientists spend thousands of micro-moments stuck in the seams between tools, formats, and workflows—doing work that’s necessary, repetitive, and oddly hard to automate well.
At Deep Origin, we’ve taken two approaches to AI development, one of which is deliberately down-to-earth: use AI where it earns its keep by removing specific, high-friction bottlenecks—especially the ones that feel mundane. Two examples:
- DO Patent extracts chemical structures from PDFs into usable digital representations (e.g., SMILES), so researchers don’t spend their week re-drawing molecules before they can do any real science.
- Why does this matter for those not in the life sciences?
- Because before most drug discovery projects can even begin, chemists often have to curate their own data — pulling molecular structures out of papers, patents, and slide decks where they exist only as images. That curation step routinely means manually re-drawing molecules by hand just to make them usable for modeling or analysis.
- Balto, our conversational interface for molecular modeling and simulation, removes the “you must code / you must learn five interfaces” tax that blocks many scientists from running serious computation, even when the underlying models already exist.
- Why does this matter for those not in drug discovery?
- Modern drug discovery relies heavily on molecular simulation — docking, scoring, and property prediction — but these tools often require months to learn, comfort with scripting, and fluency across multiple disconnected interfaces.
We also build AI in places where quality is existential—like docking and virtual screening—because if you’re going to automate a decision that drives lab spend, you’d better be right often enough to trust. We’re working on a huge moonshot through an ARPA-H award to help replace animal testing with simulations. But what’s surprised many teams we talk to is this: the fastest ROI frequently comes from the “small” problems.
This post is about why.
A little background: drug discovery is a hard place for AI (and why that matters)
From the outside, drug discovery can look like an obvious AI win.
It’s data-rich.
It’s high-cost.
It’s full of optimization problems.
So why hasn’t AI already “solved” it?
The short answer is that drug discovery is not one problem — it’s a chain of fragile decisions, each made under uncertainty, where errors compound slowly and expensively.
A single drug program typically spans:
- target identification and biological validation
- hit discovery and optimization across chemistry, potency, selectivity, and safety
- years of preclinical testing
- multi-phase clinical trials
Timelines routinely stretch 10–15 years. Costs run into the billions. And even with all that effort, the vast majority of programs fail — often late, and often for reasons that were invisible early on.
For AI practitioners, three structural realities make this domain uniquely difficult:
- Ground truth arrives late (or never)
In many AI applications, feedback loops are fast. You can A/B test. You can retrain weekly. You know quickly whether a model worked.
In drug discovery, the “label” might arrive five years later — when a molecule fails in the clinic due to toxicity, poor exposure, or lack of efficacy in humans. By then, millions of dollars and thousands of decisions have already been made.
This makes naïve end-to-end optimization not just hard, but dangerous.
- Data is fragmented, biased, and context-heavy
Unlike consumer or enterprise domains, drug discovery data is:
- sparse in the regimes that matter most (novel targets, new chemistry)
- biased toward what worked well enough to publish
- scattered across papers, patents, internal reports, and PDFs
Much of the most valuable information isn’t in clean tables — it’s embedded in figures, captions, supplementary material, or institutional memory.
- Wrong answers are far more costly than slow ones
In many AI systems, a slightly wrong output is tolerable.
In drug discovery:
- a false positive can send teams down a dead-end for months
- a false negative can kill a molecule that might have helped patients
- a poorly understood model can erode trust fast
As a result, adoption hinges not just on accuracy, but on interpretability, traceability, and workflow fit.
This context matters because it explains why progress in AI-driven drug discovery rarely comes from a single sweeping model — and why so much value is locked up in what look like “small” problems.
Why small bottlenecks dominate outcomes
Because discovery pipelines are long and interdependent, friction anywhere propagates everywhere.
If:
- chemical structures are locked in PDFs,
- only a few specialists can run simulations,
or models are powerful but inaccessible,
then even strong AI struggles to make an impact.
That’s why many of the highest-ROI applications of AI in drug discovery aren’t flashy breakthroughs — they’re workflow accelerators:
- removing translation layers between humans and machines
- compressing setup time
- reducing manual rework
- making advanced tools usable by more people, more often
This is the context in which tools like DO Patent and Balto exist — and why they matter far more than their surface simplicity might suggest.
Drug discovery doesn’t just have hard problems. It has hidden ones.
If you ask a medicinal chemist what slows a project down, you’ll hear the big themes: target biology uncertainty, ADMET surprises, synthesis complexity, translational risk.
But sit next to a team for a week and you’ll see a different class of bottleneck:
- chasing structures buried in PDF figures (patents, papers, slide decks)
- re-drawing molecules into ChemDraw
- copying structures between tools that don’t quite agree
- manually constructing datasets from documents before modeling can even begin
- waiting on the one person who knows how to run a particular workflow
- “I could do that simulation… if I remembered the flags / environment / file formats”
These aren’t glamorous problems, but they’re measurable, common, and expensive.
Even the broader data world has a well-known pattern: a large fraction of time goes to preparation and wrangling rather than “the interesting part.” An EMA data-analytics report cites a survey finding data scientists spend around 80% of their time preparing and managing data for analysis. While “data science” isn’t identical to “drug discovery R&D,” the shape of the problem is familiar to any computational group embedded in a lab: the pipeline is only as fast as its least-automated seam.
DO Patent: turning “hours of redrawing” into “minutes of extraction”
Patents and papers are full of chemistry—but much of it is effectively locked in image form. Extracting it is a known pain point in cheminformatics: the literature repeatedly describes chemical structure extraction and patent mining as difficult, time-consuming, and error-prone when done manually. ScienceDirect+1
And this isn’t a new pain. Chemists have spent enormous time simply preparing structures for communication. C&EN has described chemical structure preparation as a task someone would do “for up to four hours at a time” at a drafting table in the pre-digital era. The medium changed; the tax didn’t disappear—it often just moved into new workflows (screenshots, to redraws, to copy/paste, to reformatting, to error-checking. Then repeat.).
Why it matters:
- Time cost: If a chemist spends even 2–5 hours/week re-drawing structures from documents, that’s 100–250 hours/year per person.
- Error cost: One transposed bond or missed stereocenter can quietly poison downstream analysis.
- Opportunity cost: Those hours are taken from hypothesis generation, design, and interpretation—the work only humans can do.
The broader ecosystem is now large enough to measure the scale of the problem. For example, the PatCID paper describes 81M chemical-structure images and 14M unique structures extracted from patents—an illustration of how much chemically relevant information exists in image form. And benchmark work comparing optical chemical structure recognition (OCSR) tools emphasizes that patents are a high-stakes domain where precision and recall matter.
Our bet with DO Patent is simple: if you remove the redraw step, you collapse a recurring multi-hour workflow into minutes—while improving traceability back to the original figure (so users can validate, not just trust).
A practical ROI lens (a template you can steal)
To keep ROI honest, we like “per scientist, per week” math—because that’s where adoption lives.
Use a conservative baseline:
- Median chemist salary in the U.S. is reported around $115,000 (2024) in the ACS salary survey.
- Fully-loaded cost is commonly estimated as a multiplier on salary (benefits + overhead). Many HR/finance references cite ~1.25–1.4× as a typical range.
If a chemist costs, say, ~$115k × 1.3 ≈ $150k/year fully loaded, that’s about $75/hour assuming ~2,000 working hours/year.
Now assume DO Patent saves:
- 2 hours/week per chemist (conservative for many teams doing competitive intel or heavy literature work)
That’s:
- 2 × 52 = 104 hours/year
- 104 × $75 ≈ $7,800/year per chemist
At 10 chemists, you’re at ~$78k/year in time value recovered—before you count reduced errors, faster cycles, or the compounding benefit of better-structured internal knowledge.
Is every recovered hour “cash saved”? Not directly. But it is throughput regained, and in drug discovery throughput is often the only resource you can’t buy fast enough.
Balto: a conversational interface as an acceleration layer for modeling
A second unsexy bottleneck: access.
Most large organizations have world-class modeling tools—plus a steep gradient between “people who can operate them” and “people who need the results.” In many groups, simulation becomes a service desk: request → queue → translation → run → interpretation → back-and-forth.
Balto is designed around a pragmatic idea: make advanced workflows accessible without forcing every user to become a toolchain expert.
This isn’t “AI that discovers drugs by itself.” It’s AI that reduces:
- interface switching
- scripting friction
- hidden tribal knowledge (“the way we run this here”)
- setup overhead (formats, parameters, environment)
Why this approach works:
- Speed: the fastest way to run more modeling isn’t always a faster GPU—it’s fewer blocked humans.
- Consistency: translating intent into repeatable workflows reduces variance.
- Learning curve compression: junior scientists ramp faster; senior scientists spend less time teaching the same operational steps.
And it fits a broader industry truth: many of the biggest AI value pools in pharma are productivity-driven rather than “one model to rule them all.” McKinsey has estimated generative AI could create $60B–$110B/year in economic value for pharma and medical products, largely via productivity gains across the value chain.
Balto is our “local” version of that thesis: win back time by lowering the activation energy of computation.
Where we don’t stay down-to-earth: simulation quality
There’s a reason many AI-for-drug-discovery announcements feel like moonshots: the scientific frontier is legitimately hard. Molecular simulation is a great example—if your scoring or pose prediction is wrong, the downstream cost is paid in synthesis, assays, and months of effort.
So yes: we invest in modeling accuracy where the cost of being wrong is enormous.
But we’ve learned something important: quality AI and unsexy AI aren’t competitors. They reinforce each other.
- If docking is strong but workflows are inaccessible, you underutilize it.
- If workflows are accessible but results are unreliable, you waste lab cycles faster.
The product strategy is to build both:
- trustworthy core engines, and
- interfaces + extraction layers that remove friction around them.
Why this down-to-earth AI strategy wins (and scales)
1) It’s easier to validate
A “redraw this molecule” workflow has obvious before/after metrics:
- time-to-SMILES
- error rate
- traceability
- throughput
The tighter the feedback loop, the faster you improve the product.
2) It compounds
Saving 90 minutes once is nice. Saving 30 minutes every week across a team becomes budget, headcount flexibility, and faster iteration.
3) It aligns with how science actually ships
Drug discovery is an ensemble sport. The bottleneck moves. Teams that win are the ones that continuously remove friction across the pipeline, not the ones that bet the company on a single “breakthrough model.”
4) It’s how AI earns trust in regulated, high-stakes environments
In pharma, credibility is built by tools that work reliably on Tuesday afternoon—not just in a benchmark plot.
A closing thought: the future is not just smarter models—it’s less wasted expertise
Drug discovery will absolutely benefit from frontier AI. But in practice, the near-term advantage often goes to organizations that treat AI like an engineer treats latency: identify the hotspots, measure them, remove them, repeat.
At Deep Origin, that’s the philosophy behind DO Patent and Balto:
- make the invisible bottlenecks visible
- automate the steps nobody brags about
- reserve human attention for the work only humans can do
If the last decade taught us anything, it’s that “revolution” is usually the accumulation of many unglamorous wins—stacked until the whole system feels different.
And that’s exactly the kind of AI we like building.
About Merrill Cook
Merrill Cook is director of marketing at Deep Origin. He draws on experi Byline for Silicon Valleys Journal Attached ence across a range of marketing disciplines from paid, brand, product marketing, to content, as well as operations and web development. In the past he’s worked for or consulted with 7 seed through series B startups, often as the first marketing hire. He’s also worked as a front end developer, a data journalist, and likes to dabble in woodworking.





